Amid Avian Influenza Outbreak, Genomic Analysis of Wild Birds is Even More Important
The latest outbreak of H5N1 avian influenza has been going on in the U.S. since 2022. It has had huge consequences, with the culling of nearly 200 million birds at poultry farms and infections that have spread to many other types of animals along with dozens of humans. The outbreak is also rampant among wild birds.
Wild birds often spread avian influenza to domestic birds, so we are taking the opportunity to look back at a really informative webinar with Philip Lavretsky, associate professor at the University of Texas at El Paso, whose research is designed to bring genomic advances to the study of wildfowl. For this talk, he focused on mallards and the long-term effects of 20th century conservation efforts aimed at helping these populations. (Spoiler alert: some of those efforts may ultimately wind up doing more harm than good.)
Lavretsky aims to study as many mallard samples as possible, so he has recruited an interesting group to help: duck hunters. People who sign up for the duckDNA program can send in tissue samples and photos of the mallards they catch; Lavretsky’s team supplies free, easy-to-use submission kits. They’ve already had thousands of applicants for the program.
Those and many other samples from mallards all over the U.S. and around the world are helping Lavretsky’s team to characterize duck populations, and specifically to identify all-wild birds, domestic mallards, and hybrids, which may have breeding limitations and other survival challenges. Already, the group has found that there are far more hybrid mallards than expected, and that populations once boosted by the decades-long release of domestic mallards are now shrinking once more.
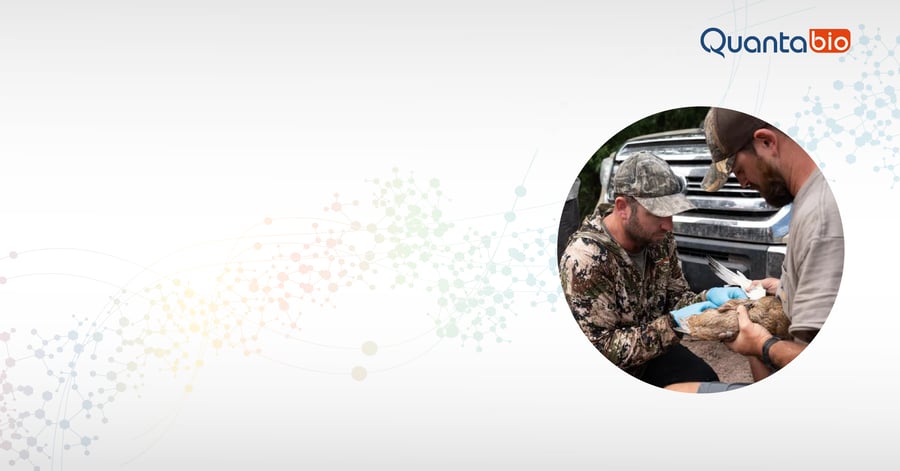
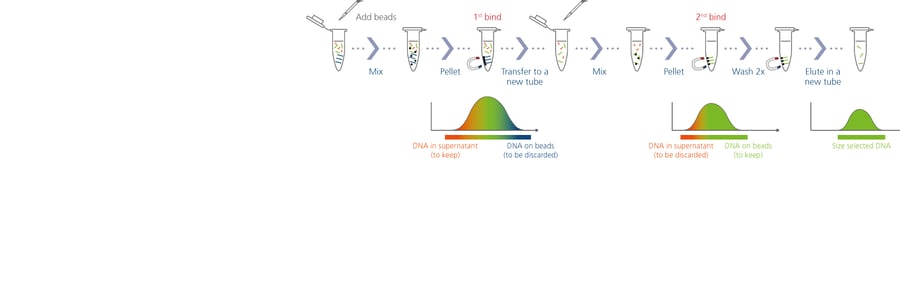
Much of the genomic analysis performed in the Lavretsky lab has come from a technique known as ddRADseq, short for double digest restriction-site associated DNA sequencing. ddRADseq is all about bang for the buck: it can provide large-scale SNP information from a huge number of genomes at very low cost compared to other types of genome sequencing. For even better results, the team added a size-selection step; they use Quantabio’s sparQ PureMag Beads, a fast and reliable nucleic acid purification system, to streamline size selection and PCR cleanup steps. The process used to take as long as 2.5 days, but now takes just 2 hours for 250 samples.
With so much genomic data from thousands of birds, Lavretsky’s team has been able to show DNA trends that indicate increases in hybrid mallards in the areas where domestic mallards were released through long-term conservation programs in the 20th century. Of course, birds migrate, and now Lavretsky says that no matter where he looks in the world, it’s possible to find hybrid mallards. Unexpectedly, his team found an area in the southern U.S. where populations were almost entirely wild; they tracked migrations and found that these mallards’ location preferences have kept them from encountering domestic and hybrid birds.
All of this might sound academic were it not for the real-world consequences. Birds with more domestic mallard DNA are less likely to migrate farther south, putting these birds at increased risk of winter weather. They were also more likely to stay in urban areas, where they might not find enough to eat. And they don’t create protective nests like wild mallards, instead dropping their eggs anywhere as farm birds might.
Going forward, Lavretsky hopes to encourage even more sample collection through the duckDNA program, which he’ll use to map hot spots of domestic mallard ancestry over time, and to hone morphology studies that will help add detail to the changes observed across domestic, wild, and hybrid birds.
To learn more about using sparQ PureMag Beads for size selection, read this application note.
Recent posts

Subscribe to Our Blog
Lorem ipsum dolor sit amet consectetur.

What Happens When Tame Meets Wild? Texas Scientists Look to Ducks for Answers
App Notes: Versatility for DNA and RNA Cleanup